Summary.
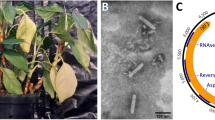
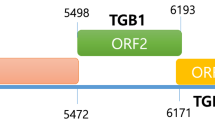
We have analysed the sequence variability in the putative reverse transcriptase (RT)/ribonuclease H (RNaseH) and the C-terminal coat protein (CP)-coding regions from Taro bacilliform virus (TaBV) isolates collected throughout the Pacific Islands. When the RT/RNaseH-coding region of 22 TaBV isolates from Fiji, French Polynesia, New Caledonia, Papua New Guinea (PNG), Samoa, Solomon Islands and Vanuatu was examined, maximum variability at the nucleotide and amino acid level was 22.9% and 13.6%, respectively. Within the CP-coding region of 13 TaBV isolates from Fiji, New Caledonia, PNG, Samoa and the Solomon Islands, maximum variability at the nucleotide and amino acid level was 30.7% and 19.5%, respectively. Phylogenetic analysis showed that TaBV isolates from the Solomon Islands showed greatest variability while those from New Caledonia and PNG showed least variability. Based on the sequences of the TaBV RT/RNaseH-coding region, we have developed a PCR-based diagnostic test that specifically detects all known TaBV isolates. Preliminary indexing has revealed that TaBV is widespread throughout Pacific Island countries. A sequence showing approximately 50% nucleotide identity to TaBV in the RT/RNaseH-coding region was also detected in all taro samples tested. The possibility that this may represent either an integrated sequence or the genome of an additional badnavirus infecting taro is discussed.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received December 11, 2002; accepted May 12, 2003
Rights and permissions
About this article
Cite this article
Yang, I., Hafner, G., Revill, P. et al. Sequence diversity of South Pacific isolates of Taro bacilliform virus and the development of a PCR-based diagnostic test. Arch Virol 148, 1957–1968 (2003). https://doi.org/10.1007/s00705-003-0163-0
Issue Date:
DOI: https://doi.org/10.1007/s00705-003-0163-0